Species Diversity¶
EcoPy contains several methods for estimating species diversity:
-
diversity(x, method='shannon', breakNA=True)¶ Calculate species diversity for every site in a site x species matrix
Parameters
- x: numpy.ndarray or pandas.DataFrame (required)
- A site x species matrix, where sites are rows and columns are species.
- method: [‘shannon’ | ‘simpson’ | ‘invSimpson’ | ‘dominance’ | ‘spRich’ | ‘even’]
shannon: Calculates Shannon’s H
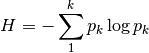
where
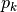 is the relative abundance of species k
is the relative abundance of species ksimpson: Calculates Simpson’s D
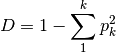
invSimpson: Inverse of Simpson’s D
dominance: Dominance index.
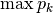
spRich: Species richness (# of non-zero columns)
even: Evenness of a site. Shannon’s H divided by log of species richness.
- breakNA: [True | False]
- Whether null values should halt the process. If False, then null values are removed from all calculations.
Example
Calculate Shannon diversity of the ‘varespec’ dataset from R:
import ecopy as ep varespec = ep.load_data('varespec') shannonH = ep.diversity(varespec, 'shannon')
-
rarefy(x, method='rarefy', size=None, breakNA=True)¶ Returns either rarefied species richness or draws a rarefaction curve for each row. Rarefied species richness is calculated based on the smallest sample (default) or allows user-specified sample sizes.
Parameters
- x: numpy.ndarray or pandas.DataFrame (required)
- A site x species matrix, where sites are rows and columns are species.
- method: [‘rarefy’ | ‘rarecurve’]
rarefy: Returns rarefied species richness.
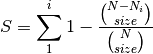
where N is the total number of individuals in the site,
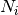 is the number of individuals of species i, and size is the sample size for rarefaction
is the number of individuals of species i, and size is the sample size for rarefactionrarecurve: Plots a rarefaction curve for each site (row). The curve is calculated as
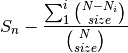
where
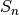 is the total number of species in the matrix and size ranges from 0 to the total number of individuals in each site.
is the total number of species in the matrix and size ranges from 0 to the total number of individuals in each site.
Example
Calculate rarefied species richness for the BCI dataset:
import ecopy as ep varespec = ep.load_data('BCI') rareRich = ep.rarefy(varespec, 'rarefy')
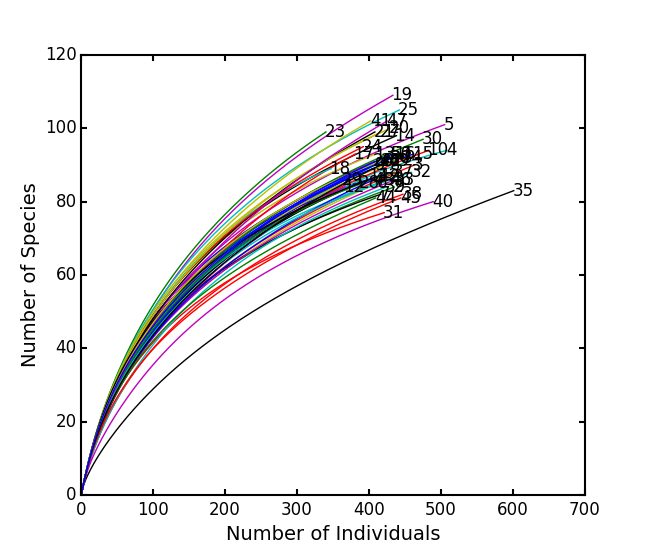
Show rarefaction curves for each site:
ep.rarefy(varespec, 'rarecurve')